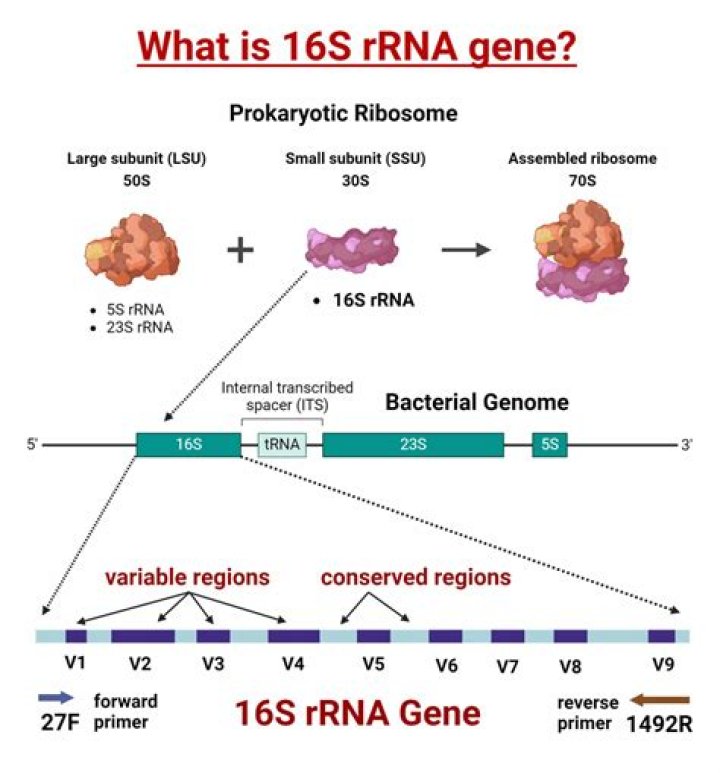
The length of 16S rRNA coding gene is about 1500bp, which contains about 50 functional domains. 16S rRNA has a number of functions: ?The immobilization of ribosomal proteins acts as scaffolding. ?3'end contains a reverse SD sequence that is used to bind to the AUG initiation codon of mRNA..
Also asked, what is the primary function of 16s rRNA?
16S ribosomal RNA (or 16S rRNA) is the component of the 30S small subunit of a prokaryotic ribosome that binds to the Shine-Dalgarno sequence. The genes coding for it are referred to as 16S rRNA gene and are used in reconstructing phylogenies, due to the slow rates of evolution of this region of the gene.
Also, what is 16s rRNA gene sequencing? 16S rRNA gene sequence analysis is a standard method in bacterial taxonomy and identification, and is based on the detection of sequence differences (polymorphisms) in the hypervariable regions of the 16S rRNA gene which is present in all bacteria.
One may also ask, why is 16s rRNA used for bacterial identification?
The 16S ribosomal RNA gene codes for the RNA component of the 30S subunit of the bacterial ribosome. Because of the complexity of DNA–DNA hybridization, 16S rRNA gene sequencing is used as a tool to identify bacteria at the species level and assist with differentiating between closely related bacterial species [8].
What is the difference between 16s rRNA and 18s RRNA?
The main difference between performing analyses with 18S rRNA gene data instead of 16S rRNA gene data (or ITS data) is the reference database used for OTU picking, the taxonomic assignments, and the template-based alignment building, since it must contain eukaryotic sequences.
Related Question Answers
Do humans have 16s rRNA?
As others have noted, 16S rRNA genes are *ubiquitous*; ribosomes can't translate mRNA without their 16S rRNA component, so all bacteria have it. Because these genes are essential, they are also very *highly conserved*. That means it is possible to construct a tree of life linking together all known bacteria.What is rRNA in biology?
Ribosomal RNA (rRNA) is part of the ribosome, or protein builders, of the cell. Ribosomes are responsible for translation, or the process our cells use to make proteins. rRNA are responsible for reading the order of amino acids and linking amino acids together. They do this through a highly complex sequence.How can 16s rRNA be used to identify bacteria?
Use of broad-range 16S rRNA gene PCR as a tool for identification of bacteria is possible because the 16S rRNA gene is present in all bacteria (Woese, 1987). The 16S rRNA gene consists of highly conserved nucleotide sequences, interspersed with variable regions that are genus- or species-specific.Why are universal 16s rDNA?
Why are universal 16S rDNA primers used in your experiment? A. They will anneal to highly conserved areas of the gene that encodes bacterial 16S rRNA. They will anneal to unique sequences of genes encoding 16S rRNA in specific bacteria.What is the difference between rRNA and rDNA?
Ribosomal DNA (rDNA) is a DNA sequence that codes for ribosomal RNA. Ribosomes are assemblies of proteins and rRNA molecules that translate mRNA molecules to produce proteins. rDNA has another gene, coding for 5S rRNA, located in the genome in most eukaryotes. 5S rDNA is also present in tandem repeats as in Drosophila.What does analysis of 16s rRNA sequences show?
In general, the comparison of the 16S rRNA gene sequences allows differentiation between organisms at the genus level across all major phyla of bacteria, in addition to classifying strains at multiple levels, including what we now call the species and subspecies level.How long is the 16s rRNA gene?
These unique DNA sequences are about 5–10 bases long and found specifically in the 16S rRNA location, and are unique to many major groups of prokaryotic organisms, archaea and Eukarya. The average lengths of the structural rRNA genes are 1,522 bp, 2,971 bp, and 120 bp respectively for 16S, 23S, and 5S rRNAs.Do eukaryotes have 16s rRNA?
The 16S rRNA gene is present in all bacteria, and a related form occurs in all cells, including those of eukaryotes.What is the 16s rRNA gene PCR used for?
BACKGROUND: Broad-range 16S ribosomal RNA (rRNA) gene polymerase chain reaction (PCR) is used for detection and identification of bacterial pathogens in clinical specimens from patients with a high suspicion for infection.What is 16s Rdna and how is it used to identify species of bacteria?
Question: What Is "16S RDNA," And How Is It Used To Identify Species Of Bacteria? 16S RDNA Is Also Called 16S RRNA DNA, Which Makes 16S RRNA To Facilitate Translation In Bacterial Cells B. 16S RDNA Is Also Called 16S RRNA DNA, Which Makes 16S RRNA To Facilitate Translation In Bacterial Cells And Eukaryotic Cells.What does the S in 16s rRNA stand for?
16S rRNA stands for 16S ribosomal ribonucleic acid (rRNA), where S (Svedberg) is a unit of measurement (sedimentation rate). This rRNA is an important constituent of the small subunit (SSU) of prokaryotic ribosomes as well as mitochondria and chloroplasts.What is metagenomic analysis?
Metagenomic analysis involves the application of bioinformatics tools to study the genetic material from environmental, uncultured microorganisms. Next generation sequencing of 16S rRNA allows the evaluation of bacterial diversity and detection of thousands of organisms.What is a universal primer?
Universal primers are complementary to nucleotide sequences that are very common in a particular set of DNA molecules and cloning vectors. Thus, they are able to bind to a wide variety of DNA templates.How many copies of 16s are in bacteria?
A total of 7,081 16S rRNA copies were identified in, with an average of 4.2 copies per genome. Fifteen percent of genomes contained a single 16S rRNA copy, while 21% contained two copies, and 3–7 16S rRNA copies per genome were frequently found.What do we mean by conserved regions in rDNA?
Conserved regions in rDNA are the sequence of nucleotides that don't change between rDNA in the organism we are studying. Comparing highly conserved regions is important to see the similarity in the DNA sequences and to examine the variable regions to see if there are any differences in structure.How are biochemical tests used to identify bacteria?
To identify bacteria, we must rely heavily on biochemical testing. Since DNA codes for protein synthesis, then different species of bacteria must, by way of their unique DNA, be able to synthesize different protein enzymes. Enzymes catalyze all the various chemical reactions of which the organism is capable.How is DNA sequenced?
DNA sequencing involves taking a DNA molecule and determining its specific sequence of nucleotides (bases). Sequencing of genomes or exomes does not involve sequencing of individual chromosomes. Instead, DNA is typically randomly fragmented into many small pieces that are each sequenced individually.Why is rRNA highly conserved?
Because rRNA is highly conserved across species through evolutionary history and is presumably present in the earliest forms of life, genes encoding rRNAs are often sequenced to determine an organism's taxonomic group, infer phylogeny (evolutionary relationships among taxonomic groups) and estimate rates of speciesWhat is shotgun cloning?
Shotgun cloning (also known as the shotgun method) is a method to duplicate genomic DNA. The DNA to be cloned is cut using a restriction enzyme or by randomly using a physical method to smash the DNA into small pieces. These fragments are then taken together and cloned into a vector.